34 Years of SoyBase
From its origins as a basic genetic map repository in the early 1990s, SoyBase has evolved into a genetics, genomics and breeding database supporting soybean research and breeding.
In the early 1990s, USDA ARS anticipated a coming genomic revolution and recognized that crop research needed a centralized, community accessible data resource. In 1992, Dr. Randy Shoemaker was selected to lead the effort to develop a soybean genetic and genomic database. At that time, SoyBase existed entirely on a single computer in the Shoemaker lab. SoyBase was developed to serve as a central repository for soybean RFLP, RAPD, SSR, and SNP marker datasets.
Data Sharing in the Pre-Internet Era

In the early days of the SoyBase, the database was distributed directly to researchers on CDs. The CDs contained an ACeDB program with the soybean genetic data. To remain current, researchers requested a CD, which the SoyBase curator mailed via USPS. Soybean researchers and breeders used the ACeDB browser on the CD to view genetic maps and query the database. The ACeDB program required the data be stored on the local computers. To remain current, researchers requested updated CDs, which the SoyBase curator mailed via USPS.
First Steps Toward Connectivity
In 1994, SoyBase was available on the World Wide Web (WWW) as a web server using an updated version of the ACeDB. Soybean researchers started citing SoyBase as the place where datasets were deposited or available via SoyBase’s FTP server—a major step forward for digital data sharing. Early scientific publications began referencing SoyBase as a source (Zhu et al., (1994), Lorenzen et al., (1995)).
At that time, SoyBase was technically “online,” but accessible only to users who knew the specific IP address. An article in Agricultural Research Magazine (1999) described this early online presence and SoyBase’s pioneering role in USDA’s efforts to provide genetic map resources over the internet.
By 2003, SoyBase needed to evolve to a platform that could both deliver data over the WWW and support remote, real-time access. SoyBase migrated to a new database management system where the data was stored in MySQL while the webpages and tools were produced with PHP. SoyBase became a scalable, maintainable, and query capable architecture, with new pages, search interfaces, and data exploration tools. These changes transformed SoyBase into a dynamic, interactive web resource accessible to researchers around the world.
Incorporating Genomic Data
SoyBase expanded dramatically in 2010 with the release of the first complete soybean genome assembly and gene model annotation from the cultivar Williams 82. This assembly—named Glyma1—represented a transformative moment for soybean research and for SoyBase. With genome-scale data available, SoyBase introduced powerful new online tools, including: GBrowse for exploring the genome and viewing gene models and markers and BLAST for sequence similarity searches. This shift marked SoyBase’s evolution from a genetic map database into a fully integrated genomic science resource.

Genomic Data Expansion within SoyBase
As high-throughput sequencing costs decreased, more genetic and genomic data was added to SoyBase. In 2014, the second and improved version of the Williams82 reference genome was released and the new soybean community gene and genome nomenclature was introduced. In 2015, the USDA soybean germplasm collection was genotyped using the SoySNP50K bead chip providing a large marker diversity data set for modern soybean breeding (Song et al., 2015).
In 2018, four new reference quality soybean genomes (Valliyodan et al., 2019; Shen et al., 2018) were released and available to view and search at SoyBase:
- Williams82 genome assembly version 4,
- Lee genome version 1,
- Zh13 genome version 1,
- and Glycine soja (PI48463) genome assembly 1.
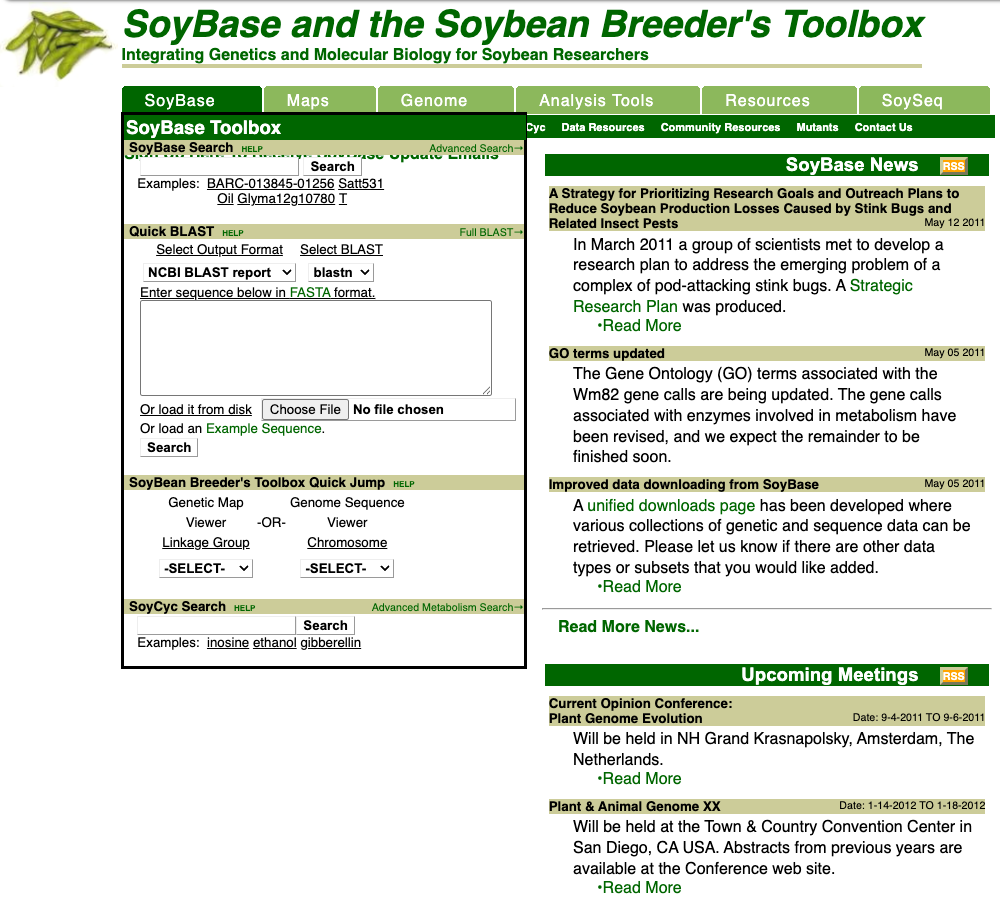
Not long after a significant number of soybean genomes were added to SoyBase, other types of genetic and genomic data were made available including the Haplotype Map for soybean GmHapMap and pan-genome data.
SoyBase started as basic genetic map and marker repository; however during the last 15 years most of the data submitted to SoyBase has been reference genomes and large-scale diversity studies. With large quantities of genomic data within SoyBase and knowing that even bigger large scale data sets would soon to arrive, the SoyBase team and major collaborators started a redesign process for both the visual look of SoyBase and the data structure.
Different look and feel but still SoyBase
From 2022 until its release in October 2024, the SoyBase team and major collaborators worked diligently to create a database that fulfilled the needs of the soybean community but also made sure the genetic and genomic data easy to find and view. During the transition the SoyBase team worked on two major tasks. The first task was to transition the data from MySQL to a new back-end database and indexed flat files. The system of flat files is referred to as the Data Store. The second task was to re-build the website architecture using updated and more secure tools. A recent publication explains in more details how the Data Store and SoyBase website architecture were built.
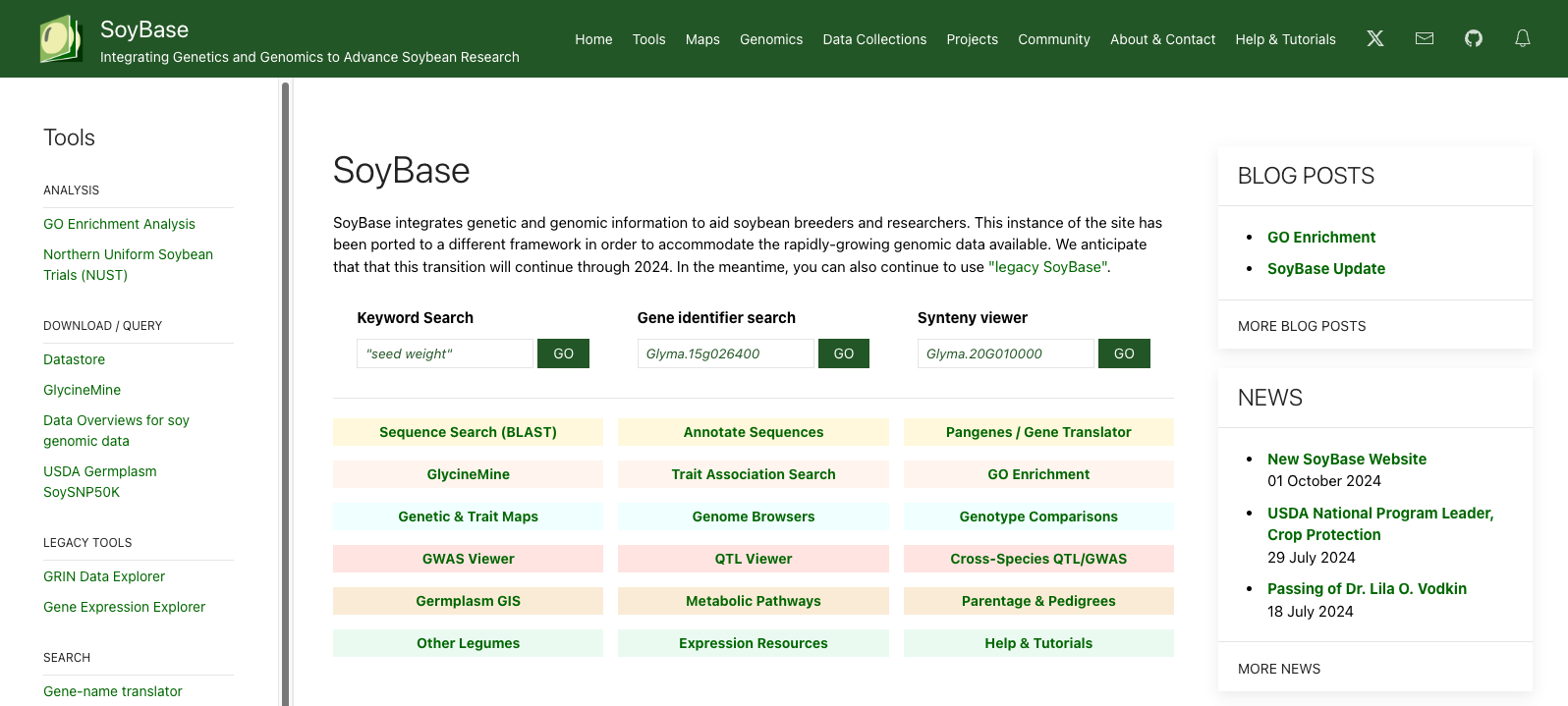
SoyBase also transitioned to different open-source tools for example:
- InterMine (GlycineMine) to house the genetic and genomic data,
- JBrowse genome browser to display genomic and genetic features in the context of a chromosome,
- and SequenceServer to allow sequence similarity searches with Blast.
At the same time, SoyBase software developer collaborators and undergraduate interns redesigned legacy tools for the new website architecture and made these tools available on GitHub for other developers. New tools for the community were created by the developers, including:
- Allele Search to search and view alleles within a variant collection,
- Gene Symbol Viewer to search gene symbols that have been published or have been submitted to SoyBase,
- Trait Association Search to search the QTL and GWAS studies within the Data Store,
- and Cmap-js to view genetic and comparative maps.
Here is to another 34 years for SoyBase
SoyBase enables breeders, researchers, and stakeholders to explore soybean genetic diversity, accelerate trait discovery, and enhance breeding efforts for resilience, nutrition, and sustainability. SoyBase ensures genetic, genomic and breeding data accessibility and fosters innovation across the agricultural research community.